Insights into mammalian TE diversity via the curation of 248 mammalian genome assemblies
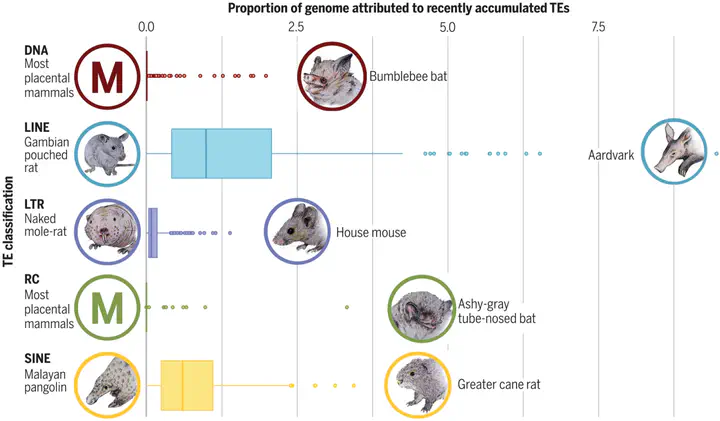 Image credit: Austin Osmanski
Image credit: Austin Osmanski
Abstract
INTRODUCTION An estimated 160 million years have passed since the first placental mammals evolved. These eutherians are categorized into 19 orders consisting of nearly 4000 extant species, with ~70% being bats or rodents. Broad, in-depth, and comparative genomic studies across Eutheria have previously been unachievable because of the lack of genomic resources. The collaboration of the Zoonomia Consortium made available hundreds of high-quality genome assemblies for comparative analysis. Our focus within the consortium was to investigate the evolution of transposable elements (TEs) among placental mammals. Using these data, we identified previously known TEs, described previously unknown TEs, and analyzed the TE distribution among multiple taxonomic levels. RATIONALE The emergence of accurate and affordable sequencing technology has propelled efforts to sequence increasingly more nonmodel mammalian genomes in the past decade. Most of these efforts have traditionally focused on genic regions searching for patterns of selection or variation in gene regulation. The common trend of ignoring or trivializing TE annotation with newly published genomes has resulted in severe lag of TE analyses, leading to extensive undiscovered TE variation. This oversight has neglected an important source of evolution because the accumulation of TEs is attributable to drastic alterations in genome architecture, including insertions, deletions, duplications, translocations, and inversions. Our approach to the Zoonomia dataset was to provide future inquirers accurate and meticulous TE curations and to describe taxonomic variation among eutherians. RESULTS We annotated the TE content of 248 mammalian genome assemblies, which yielded a library of 25,676 consensus TE sequences, 8263 of which were previously unidentified TE sequences (available at https://dfam.org). We affirmed that the largest component of a typical mammalian genome is comprised of TEs (average 45.6%). Of the 248 assemblies, the lowest genomic percentage of TEs was found in the star-nosed mole (27.6%), and the largest percentage was seen in the aardvark (74.5%), whose increase in TE accumulation drove a corresponding increase in genome size—a correlation we observed across Eutheria. The overall genomic proportions of recently accumulated TEs were roughly similar across most mammals in the dataset, with a few notable exceptions (see the figure). Diversity of recently accumulated TEs is highest among multiple families of bats, mostly driven by substantial DNA transposon activity. Our data also exhibit an increase of recently accumulated DNA transposons among carnivore lineages over their herbivorous counterparts, which suggests that diet may play a role in determining the genomic content of TEs. CONCLUSION The copious TE data provided in this work emanated from the largest comprehensive TE curation effort to date. Considering the wide-ranging effects that TEs impose on genomic architecture, these data are an important resource for future inquiries into mammalian genomics and evolution and suggest avenues for continued study of these important yet understudied genomic denizens.