Image by Laurel R. Yohe
Image by Laurel R. Yohe
Publications
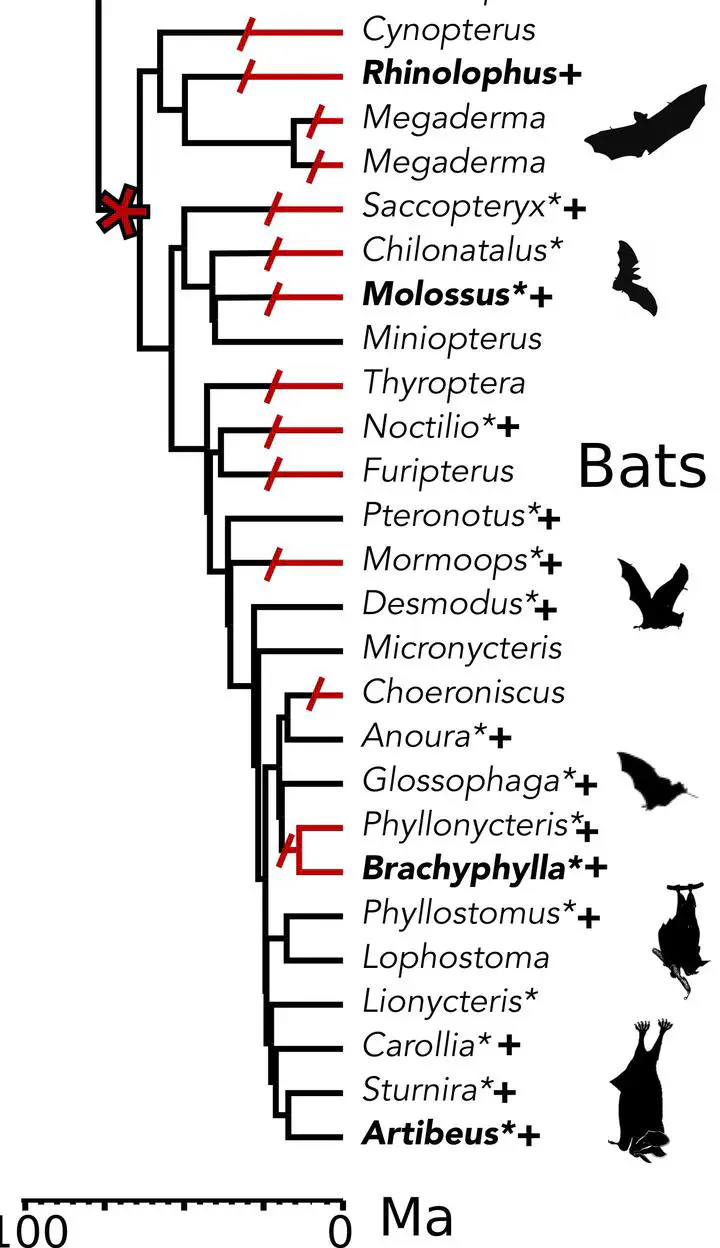
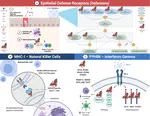
Zoonoses are infectious diseases transmitted from animals to humans. Compared to other mammalian orders, bats are suggested to harbor more zoonotic viruses(Olival et al. 2017). Infections in bats are largely asymptomatic(Schlottau et al. 2020; Guito et al. 2021), suggesting limited tissue-damaging inflammation and immunopathology. To investigate the genomic basis of disease resistance, the Bat1K project generated reference-quality genomes of ten bat species, including potential viral reservoirs. A systematic analysis covering 115 mammalian genomes revealed that signatures of selection in immune genes are more prevalent in bats compared with other mammalian orders. We found an excess of immune gene adaptations in the ancestral chiropteran branch and in many descending bat lineages, highlighting viral entry and detection factors, and regulators of antiviral and inflammatory responses. ISG15, an antiviral gene contributing to hyperinflammation during COVID-19(Perng and Lenschow 2018; Munnur et al. 2021), exhibits key residue changes in rhinolophid and hipposiderid bats. Cellular infection experiments show species-specific antiviral differences and an essential role offor protein conjugation in antiviral function of bat ISG15, separate from its role in secretion and inflammation in humans. Furthermore, in contrast to human ISG15, ISG15 of most rhinolophid and hipposiderid bats has strong anti-SARS-CoV-2 activity. Our work reveals molecular mechanisms contributing to viral tolerance and disease resistance in bats.
Ariadna E. Morales,
Yue Dong,
Thomas Brown,
Kaushal Baid,
Dimitrios - Georgios Kontopoulos,
Victoria Gonzalez,
Zixia Huang,
Alexis-Walid Ahmed,
Leon Hilgers,
Sylke Winkler,
Graham M. Hughes,
Xiaomeng Li,
Bogdan M. Kirilenko,
Paolo Devanna,
Tanya M. Lama,
Yomiran Nissan,
Martin Pippel,
Liliana M. Dávalos,
Sonja C. Vernes,
Sebastien J. Puechmaille,
Stephen J. Rossiter,
Yossi Yovel,
Joseph B. Prescott,
Andreas Kurth,
David A. Ray,
Burton K. Lim,
Eugene Myers,
Emma C. Teeling,
Arinjay Banerjee,
Aaron T. Irving,
Michael Hiller
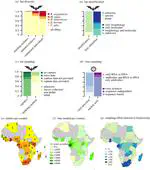
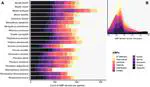
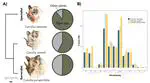
Africa experiences frequent emerging disease outbreaks among humans, with bats often proposed as zoonotic pathogen hosts. We comprehensively reviewed virus–bat findings from papers published between 1978 and 2020 to evaluate the evidence that African bats are reservoir and/or bridging hosts for viruses that cause human dis- ease. We present data from 162 papers (of 1322) with original findings on (1) numbers and species of bats sampled across bat families and the continent, (2) how bats were selected for study inclusion, (3) if bats were terminally sampled, (4) what types of ecological data, if any, were recorded and (5) which viruses were detected and with what methodology. We propose a scheme for evaluating presumed virus–host relationships by evidence type and quality, using the contrasting available evidence for Orthoebolavirus versus Ortho- marburgvirus as an example. We review the wording in abstracts and discussions of all 162 papers, identifying key framing terms, how these refer to findings, and how they might contribute to people’s beliefs about bats. We discuss the impact of scientific research communi- cation on public perception and emphasize the need for strategies that minimize human–bat conflict and support bat conservation. Finally, we make recommen- dations for best practices that will improve virological study metadata.
Natalie Weber,
Martina Nagy,
Wanda Markotter,
Juliane Schaer,
Sebastien J. Puechmaille,
Jack Sutton,
Liliana M. Dávalos,
Marie-Claire Dusabe,
Imran Ejotre,
M. Brock Fenton,
Mirjam Knörnschild,
Adrià López-Baucells,
Rodrigo A. Medellin,
Markus Metz,
Samira Mubareka,
Olivier Nsengimana,
M. Teague O’Mara,
Paul A. Racey,
Merlin Tuttle,
Innocent Twizeyimana,
Amanda Vicente-Santos,
Marco Tschapka,
Christian C. Voigt,
Martin Wikelski,
Dina K.N. Dechmann,
DeeAnn M. Reeder
Comprising more than 1,400 species, bats possess adaptations unique among mammals including powered flight, unexpected longevity, and …
Diana D. Moreno-Santillán,
Tanya M. Lama,
Yocelyn Gutiérrez Guerrero,
Alexis M. Brown,
Paul Donat,
Huabin Zhao,
Stephen Rossiter,
Laurel R. Yohe,
Joshua Potter,
Emma C. Teeling,
Sonja Vernes,
Kalina T.J. Davies,
Eugene Myers,
Graham M. Hughes,
Zixia Huang,
Federico G. Hoffmann,
Angelique P. Corthals,
David A. Ray,
Liliana M. Dávalos